  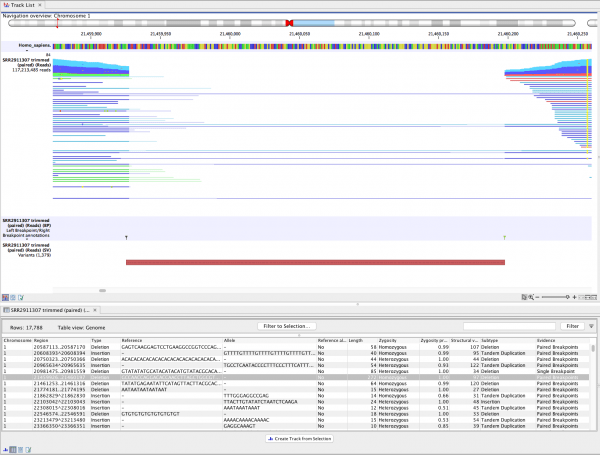 
QIAGEN Ingenuity Pathway Analysis (IPA): Deep-dive trainings – All Regions – Nov.QIAGEN OmicSoft – Powerful cloud-enabled ‘omics GUI, complete NGS analysis workflows and unparalleled curated content for immediate exploration.Human Somatic Mutation Database (HSMD) – A new somatic database developed by QIAGEN that contains extensive genomic content relevant to solid tumors and hematological malignancies.QIAGEN IPA – Powerful tools to uncover the significance of data and identify new targets or candidate biomarkers within the context of biological systems. QIAGEN Ingenuity Pathway Analysis (IPA) New user trainings – All Regions – Nov.24 – As requested by many users, QIAGEN Digital Insights team is excited to introduce QIAGEN Ingenuity Pathway Analysis (IPA) deep-dive trainings. Hereditary NGS Clinical Summit Series: Part II – Nov.23 – Join us for a 90-minute training session for new users of QIAGEN IPA. Structural Variant Detection using CLC Genomics Workbench Introduction to the Advanced Structural Variant Detection plugin for the CLC Genomics Workbench On-demand webinar: Harnessing insight from real-world oncology cases : introducing HSMD – In this on-demand webinar, we will introduce you to the Human Somatic Mutation Database (HSMD)-a new somatic database developed by QIAGEN.On-demand Webinar: How you can simplify your NGS secondary analysis workflow to 5 easy steps – Find out how you can simplify your NGS secondary analysis workflow to 5 easy steps using QCI Secondary Analysis, a new cloud-based service f.On-demand webinar: Discover hidden relationships in your toxicological studies with QIAGEN IPA – Watch this informative past webinar on how QIAGEN IPA can help you dig deeper into your toxicogenomic studies!.10 – An expert panel of leading medical geneticists, variant scientists and bioinformaticians will discuss emerging scientific and clinical trend. Structural variants affect large regions of the human genome and also play a significant role in gene expression (1, 2). They are typically detected with short Illumina or long PacBio reads, or a combination of both approaches. The new Advanced Structural Variant Detection (ASVD) plugin focuses on the short read approach, and is able to detect structural variants using short Illumina reads from whole genome sequencing (WGS). It supports the detection of the most frequently occurring structural variant types in the human genome such as deletions, duplications, and insertions (1). The ASVD plugin checks read mappings for evidence of breakpoints using “unaligned end” signatures. “Unaligned end” refers to the end of a read that does not map to the reference sequence. #Clc genomics workbench number of reads too low software#. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
